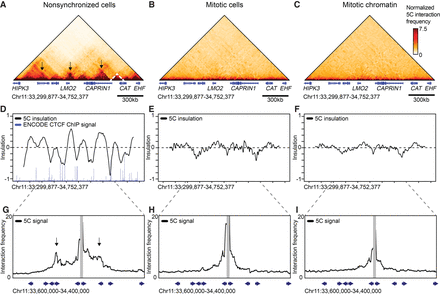
Topologically associating domains (TADs) and CTCF loops are lost in prometaphase. 5C data of Chromosome 11: 33,299,877–34,752,377 show TADs (dashed lines) and CTCF loops (arrows) in interphase (A); however, these structures are lost in nocodazole-arrested mitotic cells (B) and purified mitotic chromatin (C). (D–F) Insulation profiles. (D) The insulation profile for nonsynchronized cells shows a strong pattern alternating peaks centered within TADs and valleys at TAD boundaries. TAD boundaries are colocalized with bound CTCF as is shown by ENCODE CTCF ChIP-seq. (E,F) Insulation profiles for mitotic cells and mitotic chromatin do not show deep minima, indicating TAD boundaries are absent. (G–I) 5C interaction profiles anchored on one CTCF-bound site (15-kb bin spanning Chr 11: 34,012,377–34,027,377). (G) Peaks along these profiles (arrows) indicate CTCF loops observed in interphase. (H,I) CTCF loops are not detected in mitotic cells and mitotic chromatin. Blue arrows represent the position and orientation of CTCF motifs.











