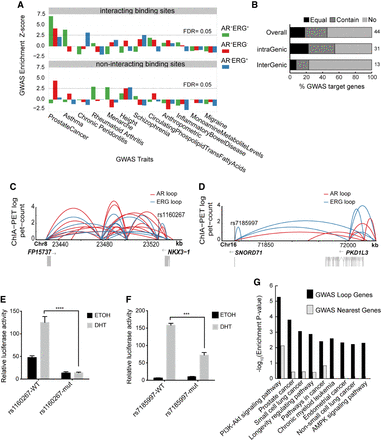
Noncoding GWAS SNPs contribute to the risk of prostate cancer through AR/ERG chromatin interaction. (A) Graph showing the enrichment of different classes of AR/ERG binding sites in diverse GWAS traits. Upper panel: Examined regions with AR/ERG chromatin looping including AR-ERG cobinding sites (green), AR only binding sites (red), and ERG only binding sites (blue) with overlapping GWAS top loci. Lower panel: Similar to the upper panel but considered only regions without AR/ERG chromatin looping. (B) The fraction of prostate cancer GWAS loci in either intra-genic regions or inter-genic regions whose targets genes (defined by AR/ERG looping) match the nearest gene: The nearest gene is the only target gene (Equal), the nearest gene is one of the target genes (Contain), and the nearest gene is not a target gene (No). (C) Prostate cancer associated GWAS SNP rs1160267 linked to the NKX3-1 gene through AR/ERG chromatin looping. (D) Prostate cancer associated GWAS SNP rs7185997 linked to PDK1L3 gene through AR/ERG chromatin looping. (E) Luciferase assay activity before and after mutating the GWAS SNP rs1160267, which is located in the enhancer region of the NKX3-1 gene. (F) Luciferase assay activity before and after mutating GWAS SNP rs7185997, which is located in the enhancer region of PDK1L3 gene. (G) A list of the top 10 pathways enriched in prostate GWAS target genes defined by either using our chromatin looping data or the nearest gene method.











