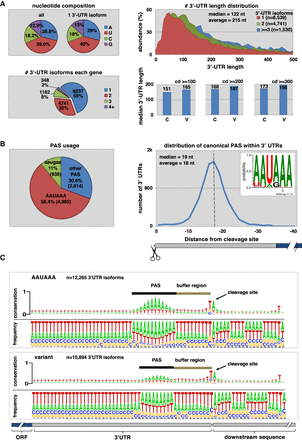
The worm 3′ UTRome v2. (A, top left) Nucleotide composition of 3′ UTRs in the 3′ UTRome v2. Uridine is the most abundant nucleotide within 3′ UTRs for C. elegans. (Bottom left) The number of 3′-UTR isoforms in each gene, and 42% of the genes in the 3′ UTRome v2 possess multiple 3′-UTR isoforms. (Top right) 3′-UTR length distribution in genes expressed with one, two, or three or more 3′-UTR isoforms. The median 3′-UTR length across these data sets is 122 nt. Genes with multiple 3′-UTR isoforms are on average longer than genes with one 3′-UTR isoform. (Bottom right) Median 3′-UTR length in genes with Canonical (C) or Variant (V) PAS elements. There is a slight increase in 3′-UTR length in genes with variant PAS elements when compared to those with canonical PAS elements. This variation is still detected when increasing the stringency of the density of the clusters (cd) used in this analysis. (B, left) PAS element usage in 3′ UTRs showing that 58.4% of 3′ UTRs use the canonical PAS element “AAUAAA,” whereas the most common variant PAS element is the hexamer “AAUGAA,” which occurs in 11% of genes. (Right) The distribution of canonical PAS elements within 3′ UTRs. The average distance from the PAS element to the cleavage site is 18 nt. (C) Alignment of 3′ UTRs at the cleavage site. This alignment in genes with both canonical and variant PAS elements reveals a region between the PAS element and the cleavage site we renamed the buffer region in which cleavage rarely occurs. The most abundant nucleotide at the cleavage site is an adenosine nucleotide preceded by a uridine nucleotide.











