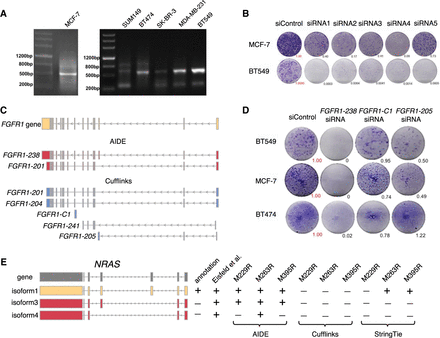
AIDE identifies isoforms with biological relevance. (A) PCR experiments validated the expression of FGFR1-238 in breast cancer cell lines MCF-7, SUM149, BT474, SK-BR-3, MDA-MB-231, and BT549. (B) Long-term colonegenic assay with Lipofectamine 3000 controls (“siControl”) and FGFR1-238 knockdowns. Tumor growths relative to the siControl were quantified by the ImageJ software (Schneider et al. 2012). (C) FGFR1 isoforms identified by AIDE and Cufflinks. (D) Long-term colonegenic assay with siControl (negative control), si-FGFR1-238 (positive control), si-FGFR1-205, and si-FGFR1-C1. Tumor growths relative to the siControl were quantified by the ImageJ software. (E) NRAS isoforms in the GENCODE annotation, reported by Eisfeld et al. (2014) and discovered by AIDE, Cufflinks, or StringTie in three melanoma BRAF inhibitor–resistant cell lines: M229R, M263R, and M395R.











