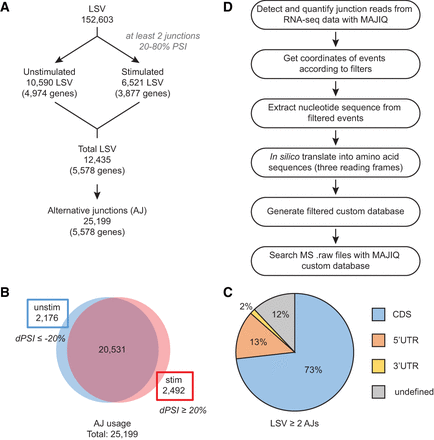
Identification of local splice variations (LSVs) and alternative junctions (AJs). (A) Workflow for identifying significantly used AJs in mRNA by identifying LSVs containing two or more exon junctions with support from 20% to 80% of the reads in either condition in previously described RNA-seq data. (B) Distribution of AJ usage in unstimulated and stimulated conditions. A shift in AJ usage is defined as the difference in percent spliced in (ΔPSI), set to 20%, between unstimulated and stimulated T cells. (C) Distribution of LSVs with two or more AJs throughout transcript regions, namely, the coding sequence (CDS) and untranslated regions (UTRs). (D) Workflow for generating MAJIQ custom protein database using AJs from A.











