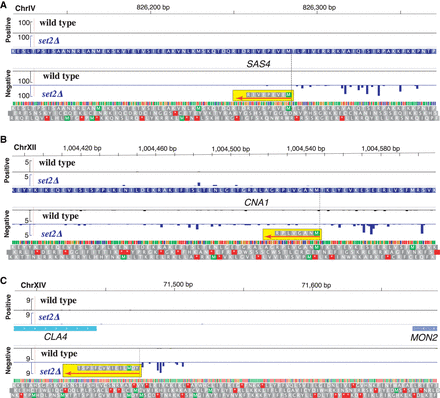
Chromatin-sensitive iTSSs encode peptides that can be detected by MS. Sequencing tracks display the 5′ cap sequence score (normalized counts) of collapsed replicates for wild type (in black) and Δset2 (in blue). Identified N-terminal peptides are highlighted in yellow, and their orientations are displayed using a red arrow. We display in gray the three potential translations of DNA in the same orientation of the detected peptide. (A) Truncation of SAS4 (MEVEPEVIR). (B) Truncation of CNA1 (MNAGVLPR). (C) Chromatin-sensitive transcript encoding a peptide in the 3′ UTR of MON2 (YDMLIEIVVCFIPST). N-terminal COFRADIC data from Varland et al. (2018).











