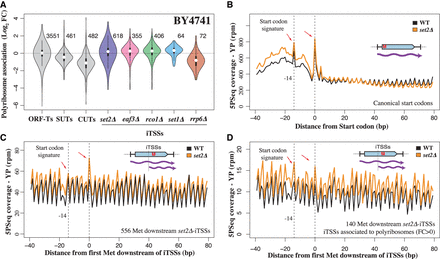
A fraction of iTSS-derived transcripts associate to ribosomes, and the internal methionine can be recognized as a novel start codon. (A) Relative association with polyribosome fraction after sucrose fractionation versus total extract. Analyzed events (present at a sufficient level in the wild-type strain) are indicated to the right of each plot. (B) Example of 5PSeq start codon–associated signature after glucose depletion for coding genes. To decrease the effect of potential outliers, we assigned a value corresponding to the 95th percentile to values that were over this threshold at each distance from the start codon. (C) Start codon–associated signature after glucose depletion for predicted novel start codons in set2Δ iTSS-derived transcripts. Those positions are expected to behave as internal methionines in a wild-type strain. (D) As in C, but showing the subset of cryptic start codons in which mRNAs are more associated with polyribosomes (fold change >0 in A).











