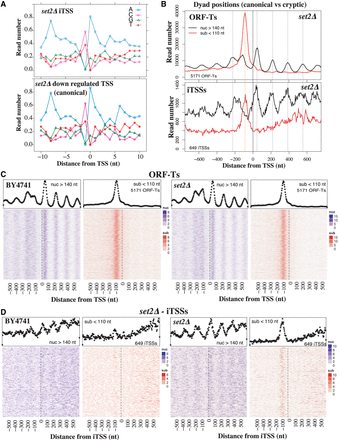
The sequence and chromatin features of iTSSs resemble those of canonical TSSs. (A) Sequence preference of set2Δ iTSSs compared with canonical TSSs (set2Δ down-regulated that often overlap with canonical TSSs). (B) MNase protection pattern for canonical ORF-T TSSs. MNase fragments are distributed in nucleosome protection fragments (nuc) and subnucleosomal ones (sub) according to their length. Vertical dotted lines depict canonical dyad nucleosome axes (in black) and putative TF binding sites (in red). (C) Heatmaps depicting in detail the MNase protection pattern for canonical ORF-T TSSs in the wild-type strain and set2Δ. Each line of the heatmaps corresponds to an analyzed region for nucleosome fragments (in blue) and subnucleosomal fragments (in red) ordered by gene expression (Xu et al. 2009). The metagene with aggregation of all the heatmap information is shown above in black dots. (D) Heatmaps depicting in detail the MNase protection pattern for set2Δ iTSSs as in C. Chromatin data are reanalyzed from Chabbert et al. (2015). Heatmap sorted by iTSS expression level.











