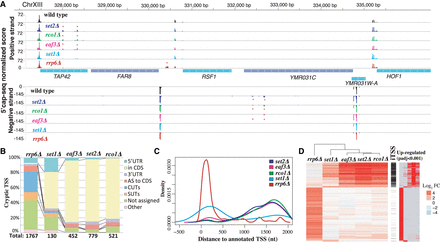
Genome-wide identification of chromatin- and RNA degradation–sensitive TSSs. Detected chromatin-sensitive cryptic transcripts tend to overlap coding genes in the same orientation. (A) Representative 5′ cap sequencing track. Score (normalized counts) of collapsed replicates is shown (see Methods). Significantly differential expressed TSSs clusters are marked by * (P-adj <0.001). (B) Classification of differentially expressed TSSs in respect to annotated features. Annotation of stable unannotated transcripts (SUTs), CUTs, and UTR lengths are from Xu et al. (2009). (C) Distribution of differentially expressed TSSs in respect to annotated ORF-T TSSs. ORF-T refer to transcripts associated with canonical ORFs as described by strand-specific tiling arrays (Xu et al. 2009). (D) Relationship between TSSs identified in the analyzed strains. Each horizontal line represents an identified TSS cluster. On the left side, we display the relative fold change enrichment (FC) with respect to the wild-type strain in log2 (red, up-regulated, to blue, down-regulated). In black, we indicate which of those identified TSSs can be classified as iTSSs. Finally, significantly differentially expressed TSSs compared to wild type are shown at the right (in red). Only TSSs identified as differentially expressed with respect to the wild type in at least one condition are shown.











