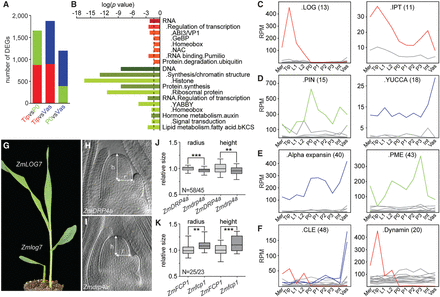
Differential expression of individual gene family members determines cell identity. (A) Pairwise differential gene expression analyses between Tip, P0, and vasculature show ∼10% of all expressed genes are differentially expressed (q < 0.01). Colors represent preferential expression within respective domains. (B) Enrichment analyses identified few functional categories overrepresented among Tip versus P0 DEGs. Dashed line, significance threshold (P = 0.05). (C–F) Individual genes within gene families show abundant and differential expression across the apex. Examples are shown for gene families functioning in cytokinin (C), auxin (D), cell wall (E), or other signaling processes (F). Number of family members is shown in parentheses. Only genes expressed ≥ 2 RPM are illustrated. Mer, meristem; Int, internode; Vas, vasculature. (G) Zmlog7 mutants display meristem termination phenotypes. (H,I) Cleared shoot apices of wild type (H) and the small meristem mutant Zmdrp4a (I). (J,K) Box-and-whisker plots of SAM size measurements of Zmdrp4a (J) or Zmfcp1 (K) and their respective wild-type siblings. (N) Number of wild-type/mutant apices measured. (**) P < 0.01, (***) P < 0.001, according to Student's t-test.











