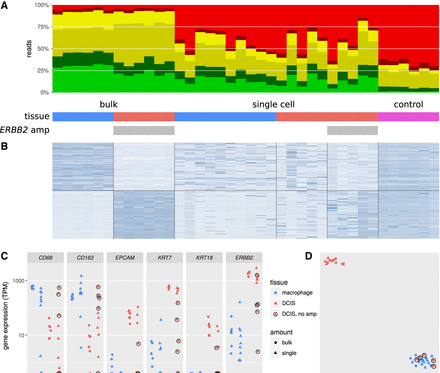
Gene expression profiling on bulk and single-cell samples from FFPE tissue dissected by LCM. (A) Alignability of Smart-3SEQ reads from FFPE bulk tissues, single cells, and negative controls. “ERBB2 amp” denotes the presence of the ERBB2 amplification in some samples (see Supplemental Fig. S5). (B) Expression (regularized log read count, normalized by row) of the 100 genes with the greatest enrichment in bulk macrophage relative to bulk DCIS and the 100 genes with the opposite enrichment, all significant at Padj < 0.01. (C) Expression (transcripts per million) of known marker genes for macrophage (CD68, CD163) and DCIS (EPCAM, KRT7, KRT18, ERBB2 [HER2]). Single cells from the DCIS tumor that lack the ERBB2 amplification are circled; we infer that these cells are intraductal macrophages. (D) t-SNE analysis of all genes; same plotting scheme as C.











