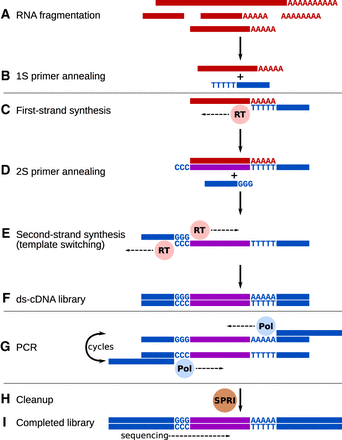
Conceptual diagram of the Smart-3SEQ library preparation method. Hands-on steps are separated by horizontal lines. (A) Total RNA is denatured and fragmented by hydrolysis. (B) The oligo(dT) primer, including a partial sequencing adapter, anneals at the beginning of the poly(A) tail. (C) Reverse transcriptase synthesizes first-strand cDNA from the RNA template and adds nontemplate dC at the end of the new strand. (D) The second primer, which includes a second partial sequencing adapter, anneals to the new dC overhang. (E) Reverse transcriptase synthesizes the second cDNA strand using the first as a template. (F) After steps C–E, which occur consecutively in one incubation, the result is a double-stranded cDNA library with partial sequencing adapters at both ends. (G) PCR with long primers amplifies the library and extends the adapters to full length, including multiplexing indexes. (H) The only cleanup step in the protocol uses paramagnetic SPRI beads to purify the amplified library while excluding adapter dimers and short inserts. (I) The final library contains the unknown cDNA sequence between the two sequencing adapters. The cDNA is sequenced in the orientation of the original RNA, yielding reads upstream of the end of the transcript (Supplemental Fig. S3). See Supplemental File 1 for a detailed technical description, Supplemental Figure S1 for a simplified diagram showing the practical workflow, and Supplemental Figure S2 for a detailed technical diagram.











