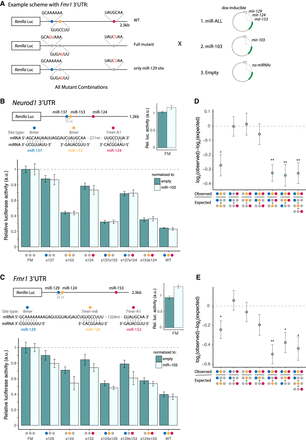
Brain cluster miRNAs collaborate to strongly repress Neurod1 and Fmr1. (A) Luciferase reporter designs (Fmr1 3′ UTR shown). Locations of seed matches to three brain cluster miRNAs are shown by colored dots, with seed and seed match sequences shown at right. (B) Neurod1 and (C) Fmr1 full-length 3′ UTRs were cloned downstream from the Renilla luciferase gene. Relative luciferase = (RmiR-ALL/FmiR-ALL)/(Rnorm/Fnorm)/FM, where RX and FX are Renilla luciferase (hRluc) and firefly luciferase (hluc+) activities in condition X, respectively. Samples were normalized to the FM from the corresponding control reporter and further normalized to either the empty plasmid (dark teal) or miR-103 control (light teal). Mean ± SD of biological triplicates is shown. (D,E) Neurod1 (D) and Fmr1 (E) observed versus expected repression from combinations of sites, as indicated under the x-axis. Error bars, SE propagated from biological triplicate measurements. Student's t-test: (*) P < 0.05, (**) P < 0.01. See also Supplemental Figure S5.











