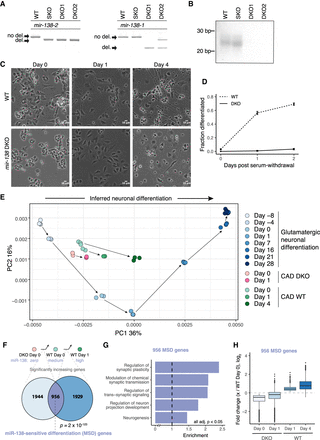
miR-138 is required for differentiation of CAD cells. (A) PCR of genomic DNA around the CRISPR-targeted sites of the two murine mir-138 loci, confirming deletion of both loci. Wild-type parental line (WT), SKO, and two DKO lines (DKO1 and DKO2) are shown. (B) Northern blot for miR-138 in the WT, SKO, DKO1, and DKO2 cell lines. (C) WT and DKO cells at 0, 1, and 4 d after serum withdrawal (10× magnification; scale bar indicates 10 µm); (D) quantitation of fraction of cells morphologically differentiated (Methods). The mean ± SD of three replicates with at least 100 cell counts each is shown. (E) PCA of RNA-seq data from eight time points of murine glutamatergic neuronal differentiation (shades of blue). RNA-seq data from CAD WT (0, 1, and 4 d after serum withdrawal; shades of green) and DKO cells (0 and 1 d after serum withdrawal; pink), projected onto PC1 and PC2 of the glutamatergic differentiation data. Arrows connect consecutive samples in each time series. The two sets of WT day 0 cells correspond to independent RNA-seq library preps and sequencing runs, as the day 0 and day 4 WT cells were sequenced in independent experiments. (F) Comparison of gene sets significantly increasing between DKO day 0 and WT day 0, and between WT day 0 and WT day 1. Significance of the 956 gene overlap (defined as MSD genes) was determined by chi-square test. (G) Gene Ontology enrichment of MSD genes against a background of genes significantly increasing in either DKO day 0 to WT day 0 or WT day 0 to WT day 1, with FDR-corrected P < 0.05 for all categories shown. (H) Fold change (log2) of the 956 MSD genes from WT day 0 to DKO day 0, DKO day 1, WT day 1, and WT day 4. See also Supplemental Figure S2 and Supplemental Tables S1 and S3.











