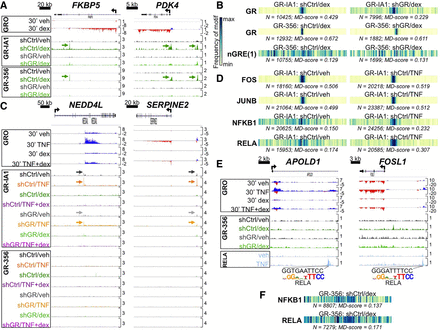
GR ChIP-seq with stable GR knockdown establishes that direct enhancer occupancy by the GR is not required for primary glucocorticoid repression of TNF targets. (A) Aligned GRO-seq and GR ChIP-seq IGV tracks at representative dex-induced loci; arrows indicate GR peaks associated with canonical GR binding sites. Vertical scales indicate maximum counts per million mapped. (B) Motif displacement (MD) analysis depicting frequency of sequence overlap with a GR and a representative nGRE binding motif within ±1500 bp of GR ChIP-seq peak summits in the indicated data sets. Each column of the barcodes is a bin of a histogram (100 bins total) where heat is proportional to the frequency of the motif instance at that distance from a ChIP-seq peak center. Darker colors signify greater enrichment on a 0–1 scale. (C) ChIP-seq and GRO-seq IGV tracks at dex-repressed loci exhibiting apparent GR occupancy in veh- and TNF-treated shCtrl cells (indicated by arrows) with the GR-IA1 but not the GR-356 antibody. (D) MD analysis of enrichment for NF-kB complex and AP-1 family motifs in the indicated ChIP-seq data sets. (E) GR-356 ChIP-seq (and GRO-seq) data combined with RELA ChIP-seq tracks (Kadiyala et al. 2016), illustrating dex-mediated repression of RELA-occupied enhancers containing NF-kB/RELA binding motifs in the absence of GR occupancy. (F) MD analysis for NF-kB complex motifs in the indicated ChIP-seq data sets.











