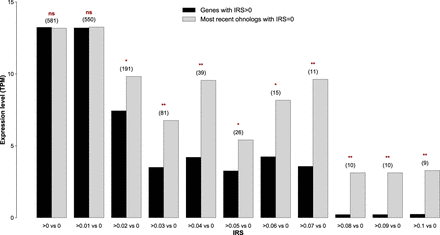
The magnitude of IES retention scales negatively with the gene expression level. The average expression level of IES-containing ohnologs (derived from the most recent whole-genome duplication) varies in an IES retention score (IRS)-dependent manner. The expression levels are based on reads mapping to the somatic genome; that is, reads from IES-containing transcripts did not go into the final estimate. Average gene expression levels were compared against one another using a one-tailed Wilcoxon rank-sum test: (ns) P > 0.05; (*) P < 0.05; (**) P < 0.01. The numbers within parentheses above the bars refer to the population size of the surveyed pairs of ohnologs.











