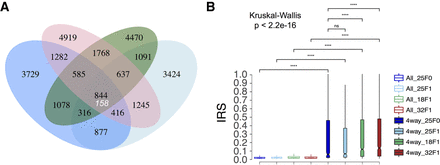
Incompletely excised IESs show elevated retention scores and might be transgenerationally inherited. (A) Venn diagram depicting sets of incompletely excised IESs (IRS > 0) shared between three F1 somatic genomes and their parental F0 genome. A large excess of incompletely excised IESs are shared across all four genomes (P < 0.001), indicating that most of the IESs in the four-way set may have been inherited transgenerationally from F0 to F1. The expected maximum number of IESs shared across all four genomes (158) (Methods) is shown under the observed value, and 229 of the observed 844 four-way-shared IESs are somatic IESs (IRS > 0.1): (blue ellipse) 25°C (F0); (green ellipse) 18°C (F1); (light blue ellipse) 25°C (F1); (red ellipse) 32°C (F1). IESs with read coverage ≥20 and length ≥26 base pairs were used. (B) Box plot showing the IRS distributions for the full set and the four-way set (4way) of incompletely excised IESs (IRS > 0) at all investigated temperatures. Pairwise comparisons between groups were performed using a Wilcoxon rank-sum test with correction for multiple testing (Benjamini–Hochberg). Statistical significance is indicated for each comparison: (****) P < 0.001; (ns) nonsignificant. Outliers are omitted for clarity.











