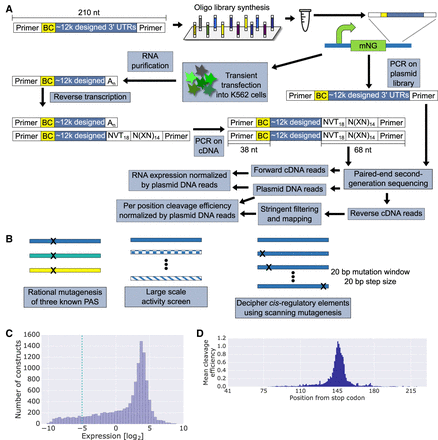
A high-throughput system for studying PAS-mediated regulation of gene expression. (A) A schematic representation of a massively parallel reporter assay measuring expression and cleavage efficiency of PAS reporters (Methods). Briefly, sequences are designed in silico, synthesized, and cloned into a reporter plasmid containing mNeonGreen. The plasmid is transiently transfected into K562 cells, from which RNA is extracted and reverse-transcribed with a poly(T) primer. N = any base synthesized randomly, V = any base except T synthesized randomly, X = A, C, G or T preselected as described in the Methods. The cDNA and plasmid DNA are amplified for paired-end second-generation sequencing. The barcodes of each library member are quantified in the forward cDNA reads and the plasmid DNA reads. These are used to calculate normalized expression. The reverse cDNA reads are mapped to their respective library members identified by the barcode in the forward DNA reads. Following stringent filtering, the cleavage efficiency distribution, normalized by the plasmid DNA reads, is calculated. (B) Library design is based on the three illustrated schemes. First, we mutated three known PASs by varying annotated regulatory elements and surrounding sequences. Second, we constructed a compendium of 6197 native PASs from annotated transcripts of viruses whose host is human and from K562 3′ end sequencing data. Finally, we applied scanning mutagenesis by mutating every 20-bp sequence in selected 629 native PASs (Methods). (C) A histogram depicting the distribution of RNA expression levels acquired by the methods in A. The −5 cutoff is used later to define positive and negative sets for motif analysis. (D) Per position mean cleavage efficiency calculated over all library variants. Positions are indicated as the distance from the mNeonGreen stop codon.











