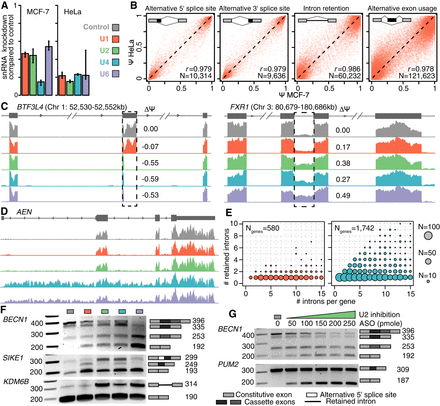
Physiological depletion of snRNAs modulates alternative splicing. (A) Levels of each snRNA following knockdown (KD) in MCF-7 and HeLa cells. Levels are relative to cells transfected with a scrambled control oligo. Error bars, 95% confidence intervals, calculated using the balanced repeated replication technique across three technical replicates of each measurement. Red indicates U1; green, U2; blue, U4; purple, U6. (B) Alternative splicing induced by U1 snRNA KD in MCF-7 versus HeLa cells. Each dot illustrates an individual splicing event (alternative 5′ splice sites, alternative 3′ splice sites, exons, and introns), represented as percent-spliced-in (PSI; Ψ) values. Plot restricted to events with at least 20 informative reads. (C) Alternative splicing of an exon within basic transcription factor 3–like 4 (BTF3L4, left) and an intron within FMR1 autosomal homolog 1 (FXR1; right). Dashed boxes indicate the differentially spliced regions. ΔΨ is defined as the fraction of transcripts containing the alternatively spliced region in snRNA KD versus control KD samples. (D) Intron retention across an entire transcript of apoptosis enhancing nuclease (AEN) following U4 or U6 snRNA KD. (E) Number of introns per gene (x-axis; plot restricted to genes with 15 or fewer introns) versus introns with a statistically significant increase in intron retention (y-axis) following KD of U1 (left) or U4 (right) snRNA. Circle size is proportional to the number of genes for each (x,y) combination. (F) PCR of select splicing events: multiple adjacent cassette exons (beclin 1, BECN1), a combination of a cassette exon and alternative 5′ splice site (suppressor of IKBKE 1, SIKE1), and intron retention (lysine demethylase 6B, KDM6B). The ladder is included on the left, and the right-hand side of each gel indicates the splice variants and their predicted sizes. (G) Dosage-dependent differential splicing (BECN1 and pumilio RNA binding family member, PUM2) following U2 KD with increasing concentration of antisense oligo (ASO).











