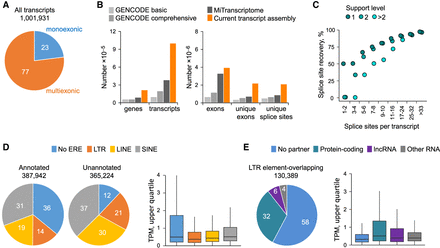
Assembly, recovery, and expression of ERE-overlapping transcripts in tumors of diverse origins. (A) Total number and proportion of monoexonic or multiexonic de novo–assembled transcripts. (B) Comparison of the total number of genes, transcripts, exons, unique exons, and unique splice sites in the current transcript assembly with GENCODE (version 24) (Frankish et al. 2019) and MiTranscriptome (Iyer et al. 2015). Genes are defined here as nonoverlapping transcribed regions. (C) Completeness of the current transcript assembly, estimated by median recovery of splice sites annotated in GENCODE. The percentage of GENCODE recovered sites is plotted according to their support levels. Recovery of the 367,411 unique splice sites of high-confidence GENCODE transcripts was ∼93%. (D) Prior annotation status and ERE composition of the 753,166 transcripts out of the entire assembly that were expressed at one or more transcripts per million (TPM) in at least one sample (left) and expression levels of these transcripts according to their ERE composition (right). Transcripts were considered as previously annotated if all exons were present within GENCODE (v24 basic) and as ERE-overlapping if any exon overlapped with an ERE integration. For transcripts overlapping with multiple EREs, we assigned a hierarchical LTR, LINE, or SINE order. As overall expression level, we used the upper quartile TPM in the cancer type with highest expression for each transcript. (E) Breakdown of LTR element–overlapping transcripts (expressed at one or more TPM in at least one sample) according to overlap with protein-coding, lncRNA, or other RNA genes (left) and expression levels (upper quartile TPM in the cancer type with highest expression) or each type of LTR element–overlapping transcript (right).











