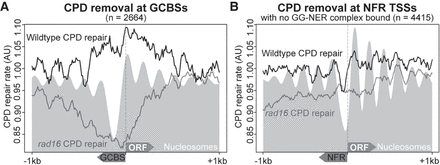
GG-NER complex binding at a subset of NFRs organizes repair in chromatin, but this is not a general feature of NFRs. (A) Relative CPD repair rates are plotted around GCBSs for both wild-type and rad16 GG-NER–defective mutant cells in relation to the nucleosome landscape, indicated as the gray shaded area. The x-axis indicates the orientation of both the GCBS and the ORF in relation to TSSs. (B) As described in A, but here plotting the relative repair rates at non-GCBS-associated NFRs (n = 4415) (see Supplemental Fig. S1), orienting the data in relation to the nearest gene aligning at the TSS with the NFR positioned upstream. The x-axis indicates regions 1 kbp upstream of and downstream from these positions. The gray shaded area represents the nucleosome data at these positions.











