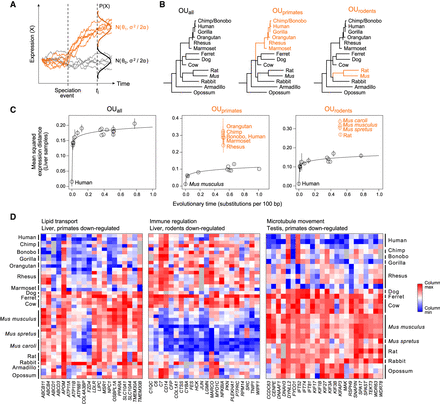
Multivariate OU process enables detection of lineage-specific expression changes. (A) Multivariate OU process. Simulated trajectories of expression (y-axis) over time (x-axis) under a multivariate OU process. Trajectories in gray are sampled from the same distribution (N0) across time, while trajectories in orange start at the same ancestral distribution (N0) but evolve under a new distribution (N1) after a speciation event. (B) Three tested hypotheses of expression evolution: from left: the univariate OUall model, in which gene expression evolves under a single stabilizing regime across the phylogeny (black), and two multivariate OU models, OUprimates and OUrodents, in which gene expression evolves under the ancestral regime (black) and a new regime in the specified subclade (orange). (C) Lineage-specific expression in liver. Pairwise mean squared expression distances in liver samples (y-axis) between a reference species (labeled black point) and each of the other mammals for genes assigned to each of three tested OU models. (Black points) species evolving under ancestral distribution; (labeled orange points) species evolving under new regime after the lineage split; (solid line) nonlinear regression fit for species evolving under ancestral distribution. (D) Example processes enriched for lineage-specific expression. Heatmaps show column-normalized expression (red: high; blue: low) from genes (columns) with lineage-specific expression patterns in three enriched GO categories (FDR < 0.05): lipid transport in liver (left), immune regulation in liver (middle), and microtubule movement in testis (right).











