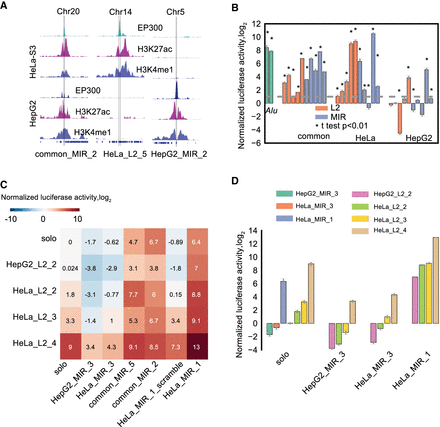
Experimental validation of the enhancer activity and interaction between MIR and L2. (A) H3K27ac, H3K4me1, and EP300 profiles for three example cases of MIR and L2 ELRs used for enhancer reporter assay. (B) Experimental validation of the enhancer activity using the luciferase reporter assay in HeLa cells. Fifteen MIR and 15 L2 ELRs were synthesized and subcloned into the luciferase reporter vector. The y-axis shows the log2 transformed normalized firefly versus Renilla luciferase activity compared with empty vector (fold change). Two Alu sequences showing the highest enhancer activity from Su et al. (2014) were used as a positive control. Each measurement was based on three biological replicates. (*) t-test P-value < 0.01 when compared to empty vector. The gray dashed line indicates the activity fold change of 1 compared to empty vector. (C) Enhancer activity for combinations of MIR and L2 using the luciferase reporter assay in HeLa cells. Mean log2-transformed activities normalized to empty vectors of three biological replicates are shown in the heat map. Each row is an L2 selected from B sorted by the activity in HeLa cells; each column is a MIR, and “solo” shows the effect of a single sequence before combination. HepG2_MIR_3 and HeLa_MIR_1_scrambled are negative controls for MIR, HepG2_L2_2 is a negative control for L2. (D) Enhancer activity measured by luciferase reporter assay in HeLa cells for the repressor and booster MIRs (labeled below the bars) on L2 elements.











