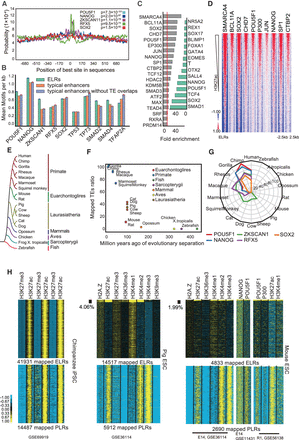
hESC- and iPSC-specific ELRs mark the master TF binding sites. (A) Top five enriched motifs detected by CentriMo around centers of ESC-specific ELRs. (B) Mean motif densities for the top 10 enriched TF motifs on ESC-specific ELRs, ESC-specific typical enhancers, and ESC-specific typical enhancer fragments that do not overlap with any TE. (C) Top enriched TFs bound at the ELRs in the ESC-specific module as determined by ChIP-seq targets, sorted by fold enrichment of TF binding over the background of all TEs. The left and right graphs are based on hESC and iPSC data from Cistrome, and data from HUES64 (GSE61475), respectively. The red line indicates the mean fold enrichment of all 67 TFs that have overlaps with tsELRs over the background of all TEs. (D) Profiles of the top 10 TFs in the left graph of panel D across −2.5 to +2.5 kb centered around tsELRs. The ELRs are sorted by the mean H3K4me1 intensity. Data sources: SMARCA4 (GSM602297), POU5F1, EP300, and NANGO, CTBP1 (H1-hESC, ENCODE), BCL11A and SP1 (GSE32465), SOX2 (GSM1364026), CHD7 (GSM831027), and JUN (GSM935614). (E) Selected representative vertebrates on the topology structure of phylogenetic tree from UCSC. (F) The mapping ratios of ELRs in the hESC-specific module in select vertebrate species. Species ages are obtained from TimeTree. (G) Radar plot of the −log10 adjusted enrichment P-values determined by CentriMo for top enriched motifs within hESC-specific ELRs in select vertebrates. (H) Histone marks and TF binding profiles in chimpanzee iPSCs (left panel, GSE69919), pig ESCs (middle panel, GSE36114), and mouse ESCs (right panel, GSE36114, GSE11431, and GSE56138) at −2.5 to +2.5 kb around mapped hESC-specific ELRs (upper panel) and PLRs (bottom panel). Only <5% and <2% of mapped hESC-specific ELRs show enhancer-like histone modifications in pig and mouse, respectively.











