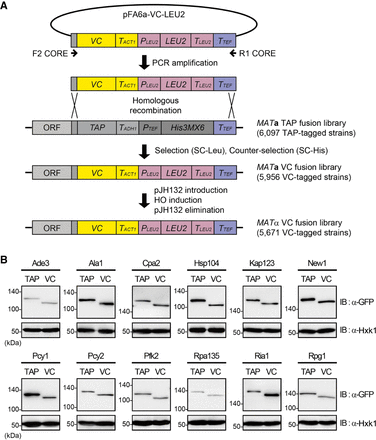
Construction of the MATα VC fusion library and confirmation of proper tagging. (A) Scheme of the construction of the MATα VC fusion library. A DNA fragment containing the VC tag and LEU2 marker sequences was amplified by PCR using the pFA6a-VC-LEU2 vector as the template and the F2CORE and R1CORE primers, and substituted for the C-terminal TAP tag sequence in the chromosome of each strain in the TAP fusion library by homologous recombination. A chromosomally VC-tagged MATα strain collection was obtained by using pJH132, as described in Methods. LEU2, PLEU2, TLEU2, TACT1, TADH1, and TTEF represent Candida glabrata LEU2, C. glabrata LEU2 promoter, C. glabrata LEU2 terminator, S. cerevisiae ACT1 terminator, S. cerevisiae ADH1 terminator, and Ashbya gossypii TEF terminator, respectively. (B) Confirmation of proper epitope switching by western blot analysis. Selected strains from the TAP fusion library and the VC fusion library were lysed and immunoblotted using an anti-GFP antibody. Hexokinase was used as a loading control. The names of proteins and the tagged epitopes are indicated at the top of each panel. The positions of molecular-weight markers are indicated on the left of each blot.











