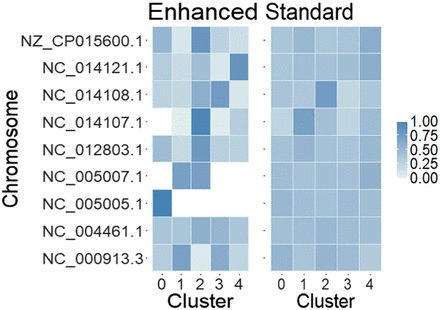
Figure 2.
Abundance of different chromosomes across clusters as assigned by Latent Dirichlet allocation (LDA). Enhanced read clouds dramatically improve LDA's ability to distinguish structure in Data set 1. This figure uses the same deconvolution as Figure 1.











