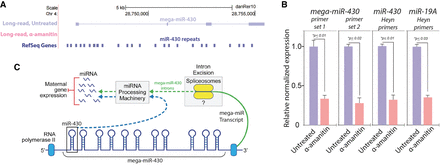
Novel validated transcript encompassing mir-430 repeats. (A) A novel transcribed region locus encompassing the mir-430 repeats. The top panel shows genomic location Chr 4: 28,745,803–28,757,055 with annotated mir-430 repeats in the reference. The two lower panels illustrate that the novel transcript found by long-read sequencing, which encompasses the mir-430 repeats, is specific to the untreated sample. (B) Experimental validation of mega-mir-430 transcript in zebrafish embryo RNA by RT-qPCR. The same RNA samples used for long-read sequencing were assayed by RT-qPCR to experimentally validate mega-mir-430. mega-mir-430 transcript was assayed using two independent primer sets spanning different introns. Both sets give nearly identical results and show susceptibility of the transcript to α-amanitin, agreeing with long-read sequencing data. Prominent early-onset “zygotic-only” genes mir-430 and mir-19a also show high abundance and strong susceptibility to α-amanitin using previously reported primers and consonant with a previous report (Heyn et al. 2014). All transcripts measured by qPCR were normalized to the hprt mRNA, observed to be unchanged during treatment. (C) Proposed functional mechanisms of mega-mir-430 transcription. mega-mir-430 serves to enrich mir-430 transcription by encoding a long single transcript that contains miRNA repeats in its introns. The generation of mature, functional miRNAs from the introns of larger transcripts is an established mechanism (Westholm and Lai 2011; Ramalingam et al. 2014). By this mechanism, mega-mir-430 gives rise to several functional miRNA units per transcript after intronic splicing and miRNA processing. If mega-mir-430 is transcribed by RNA polymerase II in parallel to polymerases generating pre-miRNAs from individual mir-430 genes, this would amplify the copies of individual miRNAs through a single larger transcript, enabling the abundant levels of mir-430 transcript observed at the start of ZGA. Illustration by Jill Gregory (Mount Sinai Health System).











