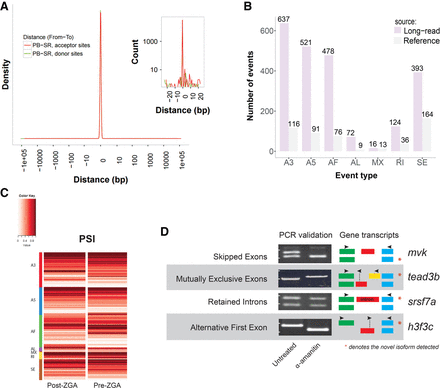
Characterization of novel isoforms for annotated gene loci. (A) Correspondence between long-read and short-read identified acceptor and donor sites in splice junctions. Shown in red is the distribution of distances from long-read-determined acceptor sites to closest short-read determined acceptor splice sites. A similar histogram is plotted for donor sites in green but is not well resolved due to superimposition of the plots. A zoom of the curves near zero is shown in the inset. Included in the plots are splice junctions for transcripts that have any overlapping introns as determined by long-read and short-read sequencing. Both curves peak heavily at zero indicating extraordinary agreement between the data sets. (B) Distribution of alternative splicing (AS) event types in the reference and long-read data. Alternative splicing events were identified using SUPPA (Alamancos et al. 2015). Listed AS includes: skipped exon (SE), alternative 5′ or 3′ splice site (A5, A3), mutually exclusive exons (MX), retained intron (RI), and alternative first or last exon (AF, AL). (C) Percent spliced in index (PSI) (Schafer et al. 2015) for novel AS events from short-read RNA-seq data in early and late embryonic stages. PSI is plotted on a zero to one scale and indicates the fraction of the total transcript amount from each locus accounted for by a specific splicing event. The PSI values corresponding to each event in panel B are plotted separately on each line and grouped by event type. For each novel AS event, PSI is shown from pre- and post-ZGA stage data. When assessed using a Wilcoxon signed-rank test, FDR-corrected q < 0.05, the data show a significant increase in alternative 3′ UTR and in retained intron usage in the late embryonic stage. (D) Validation of novel splicing events in various AS categories using PCR reactions designed to distinguish isoforms. cDNA from α-amanitin/untreated samples was used for PCR. Primer sets, indicated by arrow (→,←) pairs, were designed to flank the splicing events, and the amplicon identity was confirmed by size. Exons are represented by colored boxes. For each novel AS event, expression of two isoforms was tested in each sample. The novel isoforms directly supported by PacBio reads are indicated by (*) for each pair of isoforms shown in the “Gene transcripts” column.











