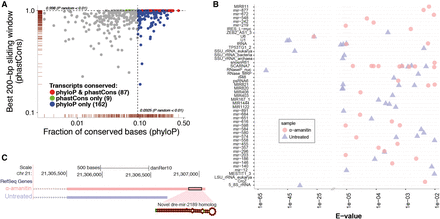
Characterization of putative noncoding NTRs. (A) Evolutionary conservation analysis of putative noncoding RNA NTRs. The scatter plot shows the base-wise transcript conservation levels (x-axis) and the maximal 200-bp window conservation levels (y-axis). Base-wise transcript conservation levels were measured as the fraction of conserved bases (base-wise phyloP score > 1.5) (Pollard et al. 2010; see Methods for score selection). Window conservation levels were measured using a sliding window average phastCons (Siepel et al. 2005) score across each 200-bp region along the transcript. Blue points indicate transcripts that showed higher base-wise conservation (phyloP) relative to randomly selected inter-genic regions (Prandom < 0.01). Similarly, green points indicate transcripts with higher window-based conservation (phastCons) relative to randomly selected inter-genic regions (Prandom < 0.01). Transcripts that met both conservation criteria are colored red. The marginal rugs along the x- and y-axes indicate the point density in each dimension. For clarity of presentation, data points with values of x < 1 × 10−3 were omitted from the plot. (B) Functional analysis of putative noncoding RNA NTRs. The cmscan option of the Infernal tools (Nawrocki and Eddy 2013) was used to search for NTRs that were good matches to the Rfam database (Griffiths-Jones et al. 2003). Each point represents an individual transcript. Multiple transcripts can be mapped to the same noncoding RNA family. The points shown are above a default significance e-value of 0.01. The high abundance of relatively lower significance assignments (to the right of the break in the x-axis) largely represents miRNAs which, due to their short structures, have a limited maximal alignment significance. (C) Novel putative miRNA homolog. A novel transcript, containing a match to the mir-548 Rfam profile, was observed in the α-amanitin sample. An alternative isoform of this transcript with a shorter 3′ tail is observed in the untreated sample. The predicted miRNA match is located in the extended 3′ region and appears to be a novel homolog of the existing zebrafish mir2189. The miRNA structure prediction was done using the Vienna RNA Websuite (Gruber et al. 2008).











