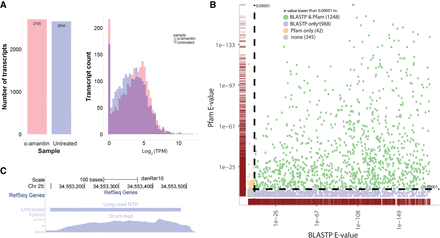
Characterization of novel transcribed regions (NTR) and coding potential. (A) Short-read support for all novel transcribed regions. Left panel: the numbers of long-read identified NTRs from the untreated and α-amanitin-treated samples. Right panel: short read support for NTRs quantified by kallisto (Bray et al. 2016). Only 270 long-read transcripts in the α-amanitin-treated condition and 343 in the untreated condition do not have short-read support (TPM < 1). Note that these NTRs include both protein-coding (see panels B and C) and non-protein-coding (see Fig. 3) transcripts. (B) Sequence homology and functional domain analysis. NTRs with high coding potential (determined by CPAT [Wang et al. 2013]) were tested for sequence homology with known proteins using BLASTP (Altschul et al. 1997) (x-axis) and for the presence of functional protein domains using hmmscan against the Pfam database (Finn et al. 2014) (y-axis). The color scheme indicates significance (e < 1 × 10−5) by both BLASTP and Pfam (green), BLASTP only (purple), or Pfam only (orange). The gray markers in the lower left corner did not achieve significance in either analysis. The point densities in each dimension are indicated by the burgundy marginal rug plots along the x- and y-axes. (C) An NTR locus with homology to the human HIST2H2BE gene. Chromosomal location Chr 25: 34,553,026–34,553,526 is shown using the UCSC Genome Browser. The empty top “RefSeq Genes” track indicates that no annotation exists in the reference. The middle track illustrates the novel transcript found by long-read sequencing in the untreated sample, and the bottom track shows corresponding mapped short reads after reference augmentation with the long-read-detected NTRs. Note the high level of expression in the short-read data.











