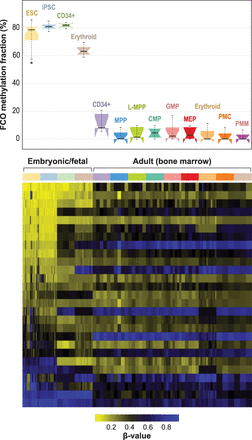
Developmentally sensitive methylation signature deconvolution in pluripotent, fetal progenitors, and adult CD34+ stem/progenitor cells. Mean (SD) estimated FCO methylation fractions for embryonic/fetal cells are 75.9% (8.5) and 4.4% (5.1) for adult progenitors (bone marrow); P = 1.81 × 10−86. In the boxplots: (1) The box shows the interquartile range (IQR), (2) the whiskers show the inner fences (1.5 × IQR out of the box), (3) the bolded line shows the median of the data, and the notches-horns display the 95% confidence interval of the median. (ESC) Embryonic stem cells; (iPSC) induced pluripotent stem cells; (CD34+ fetal) fresh cord blood cells expressing CD34+; (erythroid fetal) fetal liver CD34+ cells, differentiated ex vivo to express transferrin receptor and glycophorin; (CD34+ adult) bone marrow expressing CD34+ CD38− CD90+ CD45RA−; (MPP) multipotent progenitors; (L-MPP) lymphoid primed multipotent progenitors; (CMP) common myeloid progenitors; (GMP) granulocyte/macrophage progenitors; (MEP) megakaryocyte-erythroid progenitors; (erythroid adult) adult bone marrow CD34+ cells, differentiated ex vivo to express transferrin receptor and glycophorin; (PMC) promyelocyte/myelocyte; (PMN) metamyelocyte/band-myelocyte.











