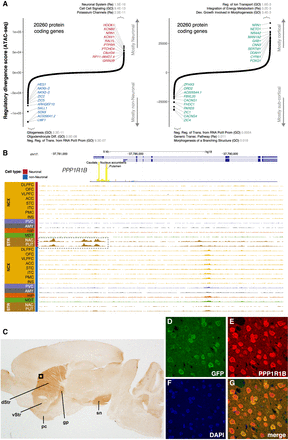
Identification of cell- and region-specific regulation of protein-coding genes. (A) Ranking of protein-coding genes based on their regulatory divergence score averaged across all neuronal versus all non-neuronal samples (left) and cortical neuronal samples versus subcortical neuronal samples (right). The regulatory divergence score is a combined measure for the difference in the regulatory burden for each gene, multiplied by how different the regulatory landscape is surrounding the gene (Methods). A gene set enrichment analysis using general gene sets and the top 500 most specific genes for either cell type/region using a one-sided Fisher's exact test was performed—the top three gene sets with P-values corrected for multiple testing using FDR are indicated. SOX8, AC009041.2, and LMF1 are all located in the same genetic locus. (B) Regional plot in the PPP1R1B locus showing OCRs. The promoter OCR and putative proximal enhancer OCRs are highlighted (dashed box). (C) The identified human PPP1R1B upstream OCR along with Exon 1, Intron 1, and the 5′ end of Exon 2 were used to direct expression of EGFP in transgenic mice. Expression identified with anti-PPP1R1B and DAB is restricted to the dorsal (dStr) and ventral striatum (vStr) (dorsal > ventral) and their projections (globus pallidus [gp] and substantia nigra [sn]) and the piriform cortex (pc). The black box indicates the region shown at higher magnification using immunofluorescence in D–G: (D) anti-EGFP (green); (E) anti-PPP1R1B (DARPP-32) (red); (F) DAPI (blue); (G) a merged image. EGFP is expressed exclusively in PPP1R1B positive neurons.











