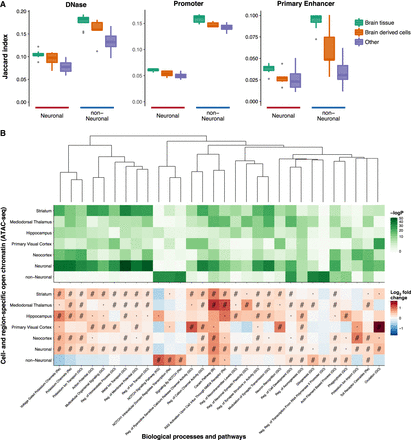
Overlap with other epigenomes and biological functions. (A) Overlap between DNase-seq OCRs and promoter/primary enhancer states of 127 epigenomes from REMC and neuronal and non-neuronal OCRs identified by ATAC-seq. Samples from REMC are split into three groups: brain tissue, brain-derived cells, and nonbrain tissues (referred to as “Other”). The full results for the individual REMC samples are shown in Supplemental Figures S11 and S12. (B) Overlap between cell- and region-specific open chromatin (ATAC-seq) and gene sets representing biological processes and pathways. Only those that were within the top five most significant gene sets in one or more ATAC-seq categories are shown. Pathways were clustered by the Jaccard index using the WardD method based on the overlap between the genes in the different gene sets and not the enrichments. This was done to show how enrichments varied by cell type and region in terms of related pathways. (#) FDR < 0.001; (·) FDR < 0.05; (Bi) BIOCARTA; (GO) gene ontology; (KG) KEGG; (Re) REACTOME. In this analysis, the region-specific OCRs were derived from neuronal samples only.











