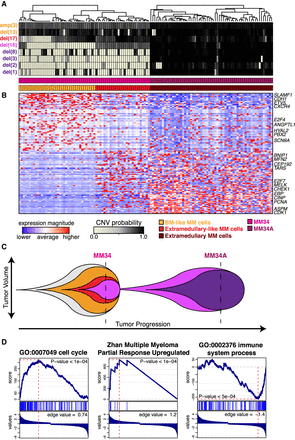
Transcriptional characterization of MM34 and MM34A. (A) Posterior probability of alterations in MM34 and MM34A. (B) Heatmap of 120 consistently significantly differentially expressed genes in comparing the BM-like MM cells versus extramedullary-like MM cells in MM34, and BM-like MM cells versus extramedullary-like MM cells in MM34 and MM34A (Supplemental Table 4). Select genes of relevance to MM or cancer based on the literature search are annotated. (C) Proposed linear pattern of subclonal evolution. (D) Gene set enrichment analysis shows enrichment in cell cycle processes (left) and a known MM partial response signature (middle) for genes up-regulated in the extramedullary-like MM34 subclone, whereas enrichment in immune response processes (right) is seen for genes up-regulated in the BM-like MM34 subclone.











