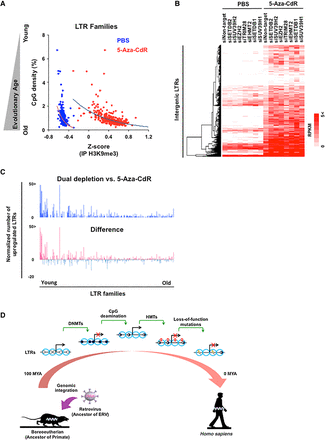
Dual depletion of DNA and histone methylation reveals an epigenetic switch in evolutionarily young LTRs. (A) The correlation between CpG density (evolutionary age) and the H3K9me3 mark in LTR families before and after 5-aza-CdR treatment. (B) The expression level of up-regulated 1033 intergenic LTRs upon dual depletion. Red indicates the expression of LTRs based on total RNA-seq. (C) The distribution of normalized number of up-regulated LTRs in each family. The bars indicate the number of up-regulated LTRs per 1000 in each family (top: dual depletion vs. 5-aza-CdR treatment; bottom: the difference between dual depletion vs. 5-aza-CdR treatment shown in the top panel and HMT KDs shown in Fig. 4C). (D) Evolutionary history of retrovirus silencing after integration into the host genome. Black dots represent DNA methylation in each CpG site, red dots represent H3K9me2/3, yellow represents DNA mutations, and blue circles represent nucleosomes. About 100 million years ago (MYA), retroviruses (ancestors of ERVs) were integrated into the boreoeutherian ancestor genome. LTR elements were silenced by DNA methylation at the beginning, but CpG sites might be lost due to CpG deamination several million years later. Therefore, the silencing mechanism switched from DNA methylation to histone methylation. Eventually, the LTR elements become silenced by the accumulation of loss-of-function genetic mutations.











