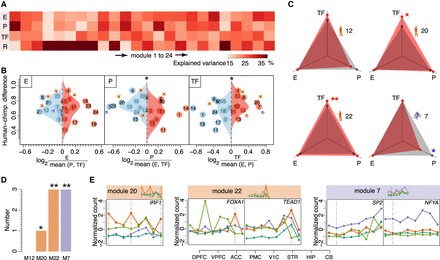
Putative spatial expression regulators. (A) The mean percentage of gene expression variation explained by epigenetic modification levels of enhancer CREs (E), promoter CREs (P), TF expression levels for TFs identified using binding sites present in proximal promoter regions (TF), and the residual variance (R) for each module. (B) Influence of the three regulatory mechanisms on spatial gene expression differences between humans and chimpanzees. The dots represent modules. The horizontal axis shows the relative influence of a regulatory mechanism in each module. The vertical axis and the distributions show human–chimpanzee expression differences for modules with the major (red) and minor (blue) contribution of a given regulatory mechanism. The asterisks indicate the significance of the difference between two distributions (one-sided Wilcoxon test, here and further: [*] P < 0.05; [**] P < 0.005). The arrows mark species-specific modules: (orange) human; (purple) chimpanzee. (C) The influence of the three regulatory mechanisms on spatial expression in species-specific modules. Red dots denote the normalized median of absolute Pearson correlation coefficients based on spatial profiles of genes and corresponding regulators. The gray dots denote values expected by chance calculated by randomly assigning regulators to genes (Methods). The asterisks represent the significance of the difference between observed and expected values (one-sided Wilcoxon test: [red] greater; [blue] smaller). (D) Numbers of TFs showing correlated expression with targets in species-specific modules. The asterisks indicate the probabilities of finding the observed or greater number of correlated TFs by chance (1000 permutations of module labels). (E) Spatial expression patterns of five TFs (IRF1, FOXA1, TEAD1, SP2, and NFYA) predicted to regulate target genes in modules 20, 22, and 7, respectively: (orange) human; (purple) chimpanzee; (dark green) gorilla; (light green) gibbon. The module expression patterns are shown above the panels. The vertical lines indicate brain regions responsible for species-specificity of the modules.











