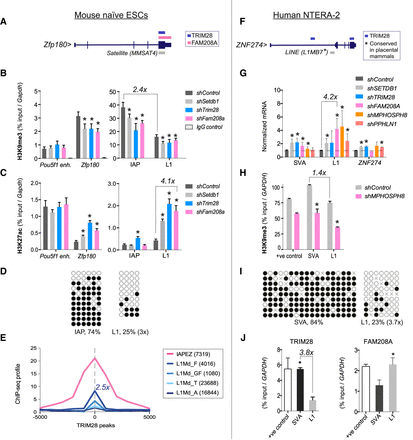
FAM208A represses L1s in leaky heterochromatin/euchromatin. (A) UCSC map of the Zfp180 locus showing TRIM28 and FAM208A binding sites and overlapping repeats (see also Supplemental Fig. S6A). (B,C) H3K9me3 and H3K27ac ChIPs on naïve mESCs. Results are representative of two (in the case of H3K9me3) or three (in the case of H3K27ac) independent IPs per treatment group performed on chromatin from the same experiment (sonicated independently), and error bars show standard deviation of all IPs, each analyzed in technical triplicates by qPCR. IgG control ChIPs gave background enrichments (ranging from 0.006 to 0.043) displayed on the H3K9me3 graph. Results are normalized to Gapdh and the Pou5f1 enhancer was used as an additional control region. Two-tailed unpaired t-tests were performed: (*) P-values <0.05. (D) DNA methylation of endogenous multicopy IAPs and L1s. (E) TRIM28 binding (Castro-Diaz et al. 2014) enrichment correlation with the stated repeat families using ChIP-cor. (F) Human ZNF274 locus showing TRIM28 binding and the presence of a conserved L1 that is bound by FAM208A in mouse cells (Fig. 5). (G) qRT-PCR of retrotransposon expression (one representative experiment of three). (H) H3K9me3 ChIP, following Mphosph8 depletion. Results are normalized to GAPDH as a negative region. +ve control; TRIM28 positive control region nearby ZNF239 (Iyengar et al. 2011). Unpaired t-tests were performed: (*) P-values <0.05. (I) DNA methylation of endogenous multicopy SVAs and L1s. (J) ChIP-PCRs using antibodies to detect TRIM28 or FAM208A binding to SVA and L1 elements. Results are representative of two independent IPs per treatment group, and error bars show standard deviation of both IPs each analyzed in technical triplicates by qPCR. IgG control ChIPs gave only background enrichment (Supplemental Fig. S7). Results are normalized to GAPDH. Positive control for TRIM28: see H; for FAM208A, we used TAF7.











