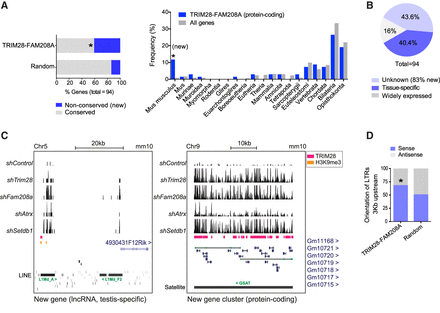
TRIM28-FAM208A coregulated genes are enriched in tissue-specific and new genes. (A, left) TRIM28-FAM208A genes and random genes were scored as conserved if they had at least 80% conservation across placental mammals (using the UCSC Table Browser). For the TRIM28-FAM208A genes, the nonconserved ones were verified to be mouse-specific using the Ensembl GeneTree. Fisher's exact test one-sided P-value = 2.204 × 10−5. (Right) Only protein-coding TRIM28-FAM208A genes (n = 68) were selected and their Last Common Ancestor extracted from the Ensembl database (version 90) using R version 3.3.1, compared to all genes in the mouse genome. Fisher's exact tests on 2 × 2 tables were significant for Mus musculus (P = 1.58 × 10−7). (B) Expression patterns of the 94 TRIM28-FAM208A genes were assessed using https://biogps.org. (C) mRNA-sequencing tracks of naïve J1 mESCs depleted of the stated epigenetic factors. TRIM28 peaks (Castro-Diaz et al. 2014) and TRIM28-dependent H3K9me3 (Rowe et al. 2013b) shown. See also Supplemental Figure S4. (D) 3 kb regions were identified upstream of each TRIM28-FAM208A coregulated gene or the random genes, and the orientation of all LTRs in these regions was assessed using the UCSC Table Browser. In the TRIM28-FAM208A group, LTRs were shown to be biased to be in a sense orientation (P = 0.005692, Fisher's exact one-sided test).











