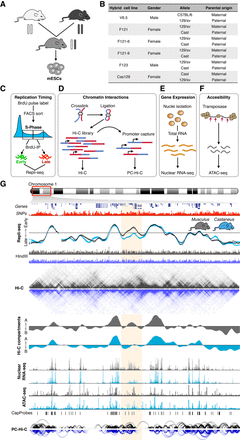
Genome-wide analysis of RT, enhancer-promoter interactions, gene expression, and chromatin accessibility in hybrid mouse ESCs. (A) Mouse ESC lines were derived from hybrid F1 blastocysts from crosses of distinct subspecies and strains. (B) Six distinct hybrid mESC lines harboring three different genomes (C57BL/6, 129/sv, and CAST/Ei), opposite parental configurations, and different genders were analyzed. V6.5, F121, and F123 hybrid cell lines were generated previously (Rideout et al. 2000; Monkhorst et al. 2008). F121-6 and F121-9 are single-cell subclones of F121, and Cas129 was generated in this study from a reciprocal cross between castaneus/musculus mice. (C–F) Genome-wide analysis of RT (C), Hi-C and promoter capture Hi-C (PC-Hi-C) (D), total nuclear RNA-seq (E), and chromatin accessibility measured by Assay for Transposase Accessible Chromatin (ATAC-seq) (F). (G) Representative genomic region on Chromosome 1 showing RT profiles of musculus (129/sv) and castaneus (CAST/Ei) alleles. Two replicates of the F121-9 cell line (two RT profiles of each genome) show the consistency in RT asynchrony. Allele-specific Hi-C matrices and compartments A and B (eigenvectors), RNA-seq, ATAC-seq, and enhancer-promoter interactions are shown. Allele-specific RT, total nuclear RNA-seq, and ATAC-seq were determined based on the SNPs shown in red. Allele-specific Hi-C and PC-Hi-C interactions were obtained using only HindIII fragments containing SNPs that distinguish each genome (HindIII track). Capture probes for PC-Hi-C are shown above promoter-enhancer interactions. Hi-C data were obtained from Giorgetti et al. (2016).











