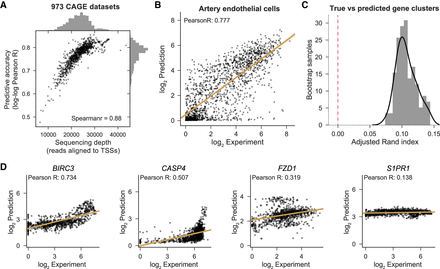
Basenji predicts cell-type–specific gene expression. (A) We computed Pearson correlation between the log2 prediction and experiment across all nonzero expressed test set genes for each CAGE data set. We plotted those correlations against the total number of reads aligned to test gene TSSs, which measures the relevant sequencing depth. (B) For the median accuracy cell, artery endothelial cells, we plotted the experiment coverage versus Basenji prediction. (C) For both the experimental measurement and Basenji prediction, the gene expression by CAGE data set matrix displays clusters. We measured the similarity of those clusters between the experimental and predicted data by bootstrap sampling gene subsets, clustering both the experimental and predicted data and computing the adjusted Rand index between the cluster sets (Methods). The adjusted Rand index is significantly greater than the null model value zero (P-value <1 × 10−26). (D) We plotted gene expression versus prediction after quantile normalization across cell types for the genes ranked in the 95th, 75th, 50th, and 25th percentiles by Pearson correlation.











