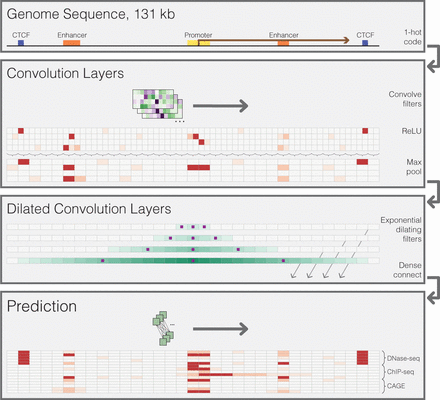
Sequential regulatory activity prediction. DNA sequences come in to the model one hot encoded to four rows representing A, C, G, and T. The annotations are fabrications to help convey the reasons for the various elements of the architecture. We apply several layers of convolution and max pooling, similar to previous methods (Kelley et al. 2016), to obtain representations that describe 128-bp bins. To share information across large distances, we apply several layers of dilated convolutions. The purple squares indicate the columns that the convolution directly sees; the teal shade is drawn proportional to the number of operations performed on that column with respect to the center position. Dilated convolution layers are densely passed on to the final prediction layer, where a width-one convolution layer makes predictions across the sequence. We compare these predictions to the experimental counts via a Poisson regression loss function and use stochastic gradient descent with back propagation to fit the model parameters.











