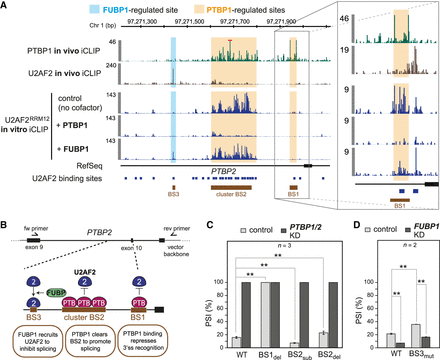
In vitro experiment captures in vivo regulation at PTBP2 exon 10. (A) In vitro cofactor assays recapitulate in vivo clearance and stabilization of U2AF2 binding by PTBP1 and FUBP1, respectively. Genome browser view of in vivo iCLIP for PTBP1 (green) and U2AF2 (brown), as well as in vitro iCLIP for U2AF2RRM12 alone and upon addition of recombinant FUBP1 and PTBP1 proteins upstream of PTBP2 exon 10. Regulated U2AF2 sites are marked as BS1-3. Red bar marks in vivo PTBP1 iCLIP signal that extends beyond the visualized range. (B) Minigene reporter assays reveal regulatory impact on alternative exon inclusion. (Top) RT-PCR primers to measure splicing changes. (Bottom) Schematic model of U2AF2 regulation at BS1-3 and its impact on inclusion of PTBP2 exon 10. (C,D) Inclusion of PTBP2 exon 10 is conjointly regulated by BS1, BS2, and BS3. Bar plots showing PTBP2 exon 10 inclusion (depicted as “percent spliced in,” PSI) in control (light gray) and PTBP1/2 (C) or FUBP1 (D) knockdown (KD, dark gray) HeLa cells. Minigene constructs include wild-type (WT) and four mutated versions with BS1 deletion (BS1del), BS2 deletion (BS2del), substitution of BS2 with a U2AF2 binding site that is not regulated by PTBP1 (BS2sub), and BS3 mutation that eliminates the U2AF2 recognition motif (BS3mut). (**) P-value < 0.01 (two-sided Student's t-test). Error bars represent standard deviation of the mean (n = 2 for FUBP1 and n = 3 for PTBP1 knockdown experiments).











