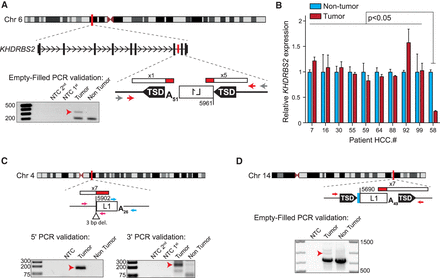
Tumor-specific L1 insertions in HCC patients with a history of alcoholism. For each insertion, the chromosomal location and orientation relative to the interrupted gene, if applicable, are shown. L1 insertions are depicted as white rectangles; the truncation point of each insertion is indicated. Target-site duplications (TSDs) are shown as black arrows; deletions of genomic DNA are shown as white triangles. Poly(A) tail length is indicated. Where applicable, untemplated nucleotides at the 5′ L1-genome junction are indicated as a blue rectangle. The location and count of junction-spanning RC-seq reads supporting each insertion are depicted as red and white rectangles. The positions of PCR validation primers are shown as small arrows. Where empty-filled validation was successful, primers are indicated in red. Where junction PCRs were employed, primers are depicted in pink (5′ junction) and blue (3′ junction). Where nested or hemi-nested PCR was necessary, the outer primer(s) are depicted in gray and the inner primers are depicted in color. Agarose gels containing the empty/filled or 5′ and 3′ junction validation products are shown. Templates are indicated above; for nested and hemi-nested PCR reactions, NTC 1st and NTC 2nd depict the no-template control reactions for the first and second rounds of PCR. Red arrows indicate the validating band. (A) A 5′ truncated L1 insertion in antisense orientation within the seventh intron of the gene KHDRBS2, detected in patient HCC.58 tumor. (B) KHDRBS2 mRNA abundance relative to TBP measured by qRT-PCR using RNA extracted from 10 HCC patient tumor (red) and nontumor (light blue) sample pairs, including patient HCC.58. Data represent the mean of three technical replicates ± SD. For each patient, values were normalized to the nontumor mean. Significance values were obtained via a two-way ANOVA followed by Tukey's post hoc analysis. (C,D) Two additional tumor-specific 5′ truncated L1 insertions, their genomic locations and structural hallmarks, and validating PCR products are depicted as described above.











