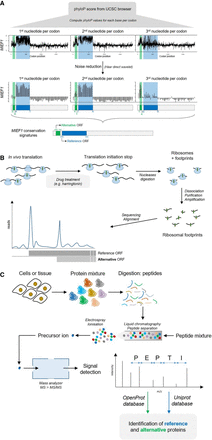
Large-scale detection methods for alternative proteins detection. (A) Conservation signatures of proteins encoded in different reading frame from the same mRNA. PhyloP scores can be computed from the UCSC Genome Browser, and noise filtration (by Haar direct wavelet) allows for the identification of distinct purifying selection signals in each reading frames. Here, the example of the dual coding MIEF1 gene is represented and corroborates data from mass spectrometry and ribosome profiling with the detection of an alternative ORF upstream of the canonical CDS (reference ORF). (B) Schematic representation of the ribosome profiling technique. This technique allows for detection of ribosomal footprints, and subsequent mapping on the genome yields a map of translation events throughout. Translation initiation at alternative ORFs can then be detected. (C) Schematic representation of the mass spectrometry technique. The search space bears crucial consequences on peptide identification. Here, we represent the strategy used by the OpenProt database that re-analyzed published mass spectrometry studies adding their predicted alternative ORFs to the scope of possibilities.











