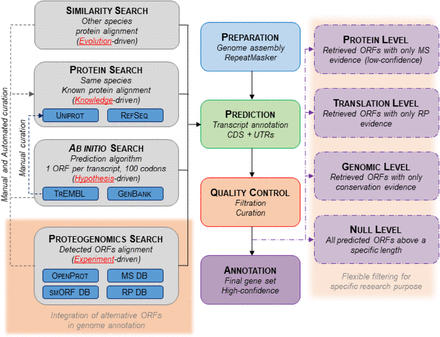
Proposed novel genome annotation framework. Current genome annotations’ pipelines have four main steps: Preparation, Prediction, Quality Control, and Annotation. The Prediction step aims to annotate transcripts (exons, introns) and CDSs (with flanking UTRs). It mostly relies on three methods: a search by homology (different species known proteins are aligned to the genome assembly), a search by prior knowledge (same species known proteins are aligned to the genome assembly), and a search ab initio (prediction of ORFs by algorithms). Here, we suggest adding an experiment-driven search and including alternative ORFs with experimental detection. The output could also be flexible to fit different experimental purposes. The pipeline steps highlighted in red correspond to the suggested implementation.











