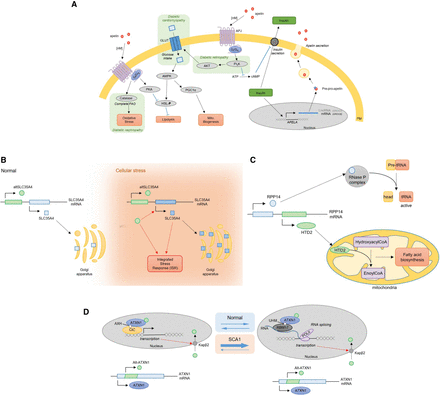
Examples of biologically important alternative ORFs. (A) Apelin, from overlooked to metabolic regulator. Apelin is encoded in an mRNA (GRCh38), previously annotated lncRNA (GRCh37), and subsequently secreted. Upon binding with APLNR (also known as APJ) receptor at a nanomolar range, it stimulates different metabolic pathways (glucose uptake, fatty acid oxidation, and mitochondrial biogenesis) and inhibits others (lipolysis and insulin secretion). These pathways are also involved in diabetic complications (cardiomyopathy, nephropathy, and retinopathy). Blue arrows represent inhibitory relationships, pathways involved in diabetic complications are highlighted in green. FAO: fatty acid oxidation; PM: plasma membrane; lncRNA: long non-coding RNA; Mito: mitochondrial. (B) SLC35A4 and its uORF-encoded protein, alt-SLC35A4. The SLC35A4 mRNA encodes two ORFs. Under physiological conditions, the canonical ORF, SLC35A4, is weakly expressed. The uORF-encoded protein alt-SLC35A4 is suspected to be the major protein product. Under cellular stress, both proteins are expressed. The alt-SLC35A4 expression level remains unchanged but positively regulates expression of SLC35A4. Both proteins are thought to be involved in the integrated stress response. ISR: integrated stress response. (C) RPP14 and its dORF-encoded protein, HTD2. The RPP14 mRNA encodes two ORFs. The canonical ORF encodes a member of the ribonuclease P (RNase P) complex (RPP14) involved in tRNAs maturation. In the 3′ UTR, a second ORF encodes a mitochondrial dehydroxylase, HTD2. HTD2 is involved in mitochondria fatty acid synthesis. (D) ATXN1 is a dual coding gene. ATXN1 mRNA encodes two proteins, ataxin and alt-ataxin. Upon entry into the nucleus, ataxin binds the transcription factor capicua (CIC) and associates with DNA at transcription sites. Ataxin nuclear localization and transcription are necessary for alt-ataxin nuclear import and its interaction with ataxin in nuclear inclusions. Ataxin is thought to shuttle between CIC complexes and RNA-binding RBM17 complexes. Polyglutamine extensions in ataxin are responsible for spinocerebellar ataxia type 1 (SCA1) and alter the dynamics of ataxin localization, thereby altering gene expression.











