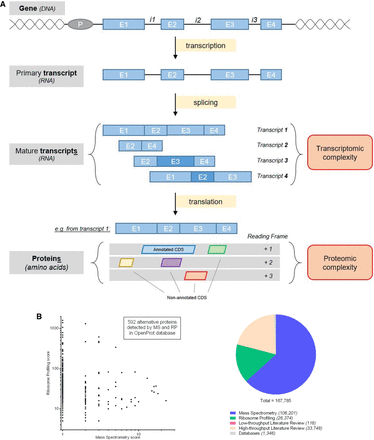
Schema of the transcriptomic and proteomic complexity inherent to a gene. (A) Genomic complexity representation. A gene is represented with a promoter (P) and introns (i) and exons (E). Splicing events lead to a suite of transcripts with frameshifted exons (darker blue shade), skipped exons, or retained introns. Then, proteomic complexity comes from each transcript with ORFs from any reading frames. However, now only one CDS is annotated per transcript, leaving an entire hidden proteome (unannotated CDSs). (B) Alternative ORFs databases. The OpenProt database predicts every ORF longer than 30 codons and reports experimental detection evidence for each of them. Five hundred ninety-two alternative ORFs were detected by both ribosome profiling (RP) and mass spectrometry (MS). The SmProt database reports smORFs (<100 codons) in different data sets (mass spectrometry, ribosome profiling, literature mining, and databases).











