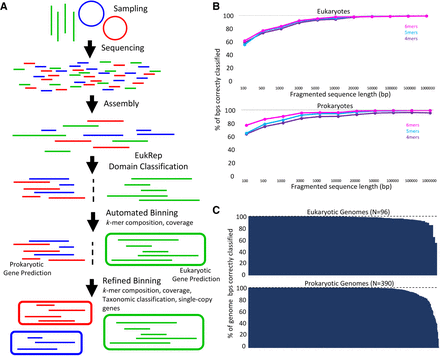
Identification of scaffolds for eukaryotic gene prediction with EukRep. (A) Schematic of the analysis pipeline used to identify and bin both eukaryotic and prokaryotic genomes within this paper. (B) A subset of genomes from Supplemental Table S2 was used to compare prediction accuracy of linear-SVM models trained on k-mer frequencies of k-mers ranging in length from 4 to 6 bp. For each sequence size category, sequences longer than the specified length were fragmented to the specified length and sequences shorter were excluded. (C) Accuracy of EukRep domain prediction on a per-genome level for both eukaryotes and prokaryotes. Percent of the genome correctly classified is defined as the percent of base pairs within a given genome predicted to belong to the genome's known domain. Each bar represents the percent of a single genome that was classified correctly. Genomes used for training and testing of EukRep along with their prediction results are listed in Supplemental Tables S1 and S2.











