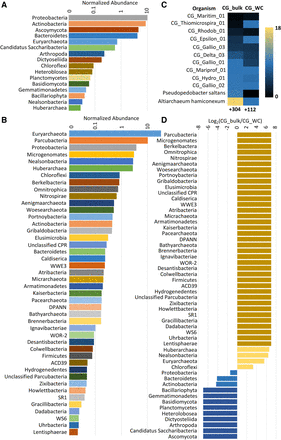
Comparison of CG_WC and CG_bulk community composition. The relative abundances of taxonomic groups in CG_WC (A) and CG_bulk (B) are depicted. Abundance was determined as the average coverage depth of the scaffolds containing annotated ribosomal protein S3 (rpS3) genes. Abundances were normalized for comparison across samples by multiplying the average coverage depth by the sample read count and read length. (C) Normalized coverage of rpS3 containing scaffolds of strains common to both samples. The number of additional strains detected in each sample is listed below the respective sample heat map. (D) Log2 ratio of normalized coverage of taxonomic groups from A and B. Taxonomic groups identified in only one sample are indicated by the darker yellow and blue bars.











