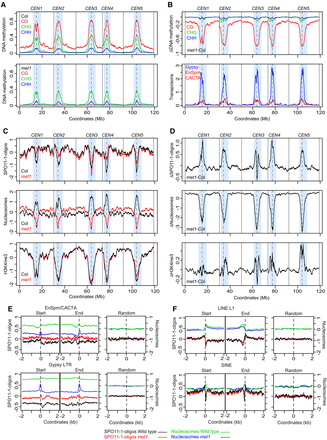
Coordinate epigenetic remodeling of chromatin and SPO11-1-oligonucleotides in met1 DNA methylation mutants. (A) Percent DNA methylation in CG (red), CHG (green), and CHH (blue) sequence contexts plotted from wild type (Col) and met1-3 (Stroud et al. 2013). Values were calculated in adjacent 10-kb windows and plotted along the A. thaliana chromosomes, using a rolling average. Centromeric assembly gaps are indicated by vertical dashed lines, and telomere positions are indicated by vertical solid lines. The pericentromeres are shaded light blue, which are defined as regions with greater than average DNA methylation. (B) The wild type (Col) versus met1 DNA methylation differential (Δ, met1 − wild type), plotted as in A (upper). The density of Gypsy (blue) and EnSpm/CACTA (red) transposons are plotted as in A (lower). (C) SPO11-1-oligos (Z-score standardized log2[SPO11-1-oligos/gDNA]), nucleosome occupancy (Z-score standardized log2[MNase/gDNA], blue), and H3K4me3 (Z-score standardized log2[ChIP/Input], blue) plotted as for A, in wild type (Col, black) and met1-3 (red). (D) The wild type (Col) versus met1 SPO11-1-oligo, nucleosome occupancy and H3K4me3 differentials (Δ, met1 − wild type), plotted as in A (upper). (E) Plots analyzing SPO11-1-oligos in wild type (black) and met1 (red), and nucleosomes in wild type (green) and met1 (blue) for EnSpm/CACTA and Gypsy transposons in 4-kb windows around their start and end coordinates, or the same number of randomly chosen positions. (F) As for E, but analyzing LINE L1 and SINE transposons.











